WEHI-539
WEHI-539 is a small-molecule inhibitor of BCL XLwith an IC50 value of 1.1 nM [1].
WEHI-539 was designed as a BCL-XLinhibitor with high affinity. It interacted the with the binding groove of BCL-XLwith a Kd value of 0.6 nM. In MEF cells lacking MCL-1, WEHI-539 induced apoptosis which was evidenced by the release of mitochondrial cytochrome cand caspase-3 processing. In BCL-XLoverexpressed MEF cells, WEHI-539 showed EC50 value of 0.48 μM. WEHI-539 also significantly induced apoptosis of the platelets purified from mice. Besides that, WEHI-539can not kill MEF cells lacking BAK because the cell death mediator BAK is regulated by BCL-XLand MCL-1. [1].
References:
[1] Lessene G, Czabotar P E, Sleebs B E, et al. Structure-guided design of a selective BCL-XL inhibitor. Nature chemical biology, 2013, 9(6): 390-397.
- 1. Iseghohimen, Adesuwa. "Overcoming drug resistance: targeting the BCL-2 family and the long non-coding RNA HCP5 in medulloblastoma and colorectal cancer." University of Salford Manchester, Jul 3, 2023.
- 2. Venneker, S. "Exploiting vulnerabilities induced by recurrent mutations in chondrosarcoma and giant cell tumour of bone: therapeutic targeting of the altered epigenome and beyond." Leiden University Scholarly Publications. Jan 01, 2023.
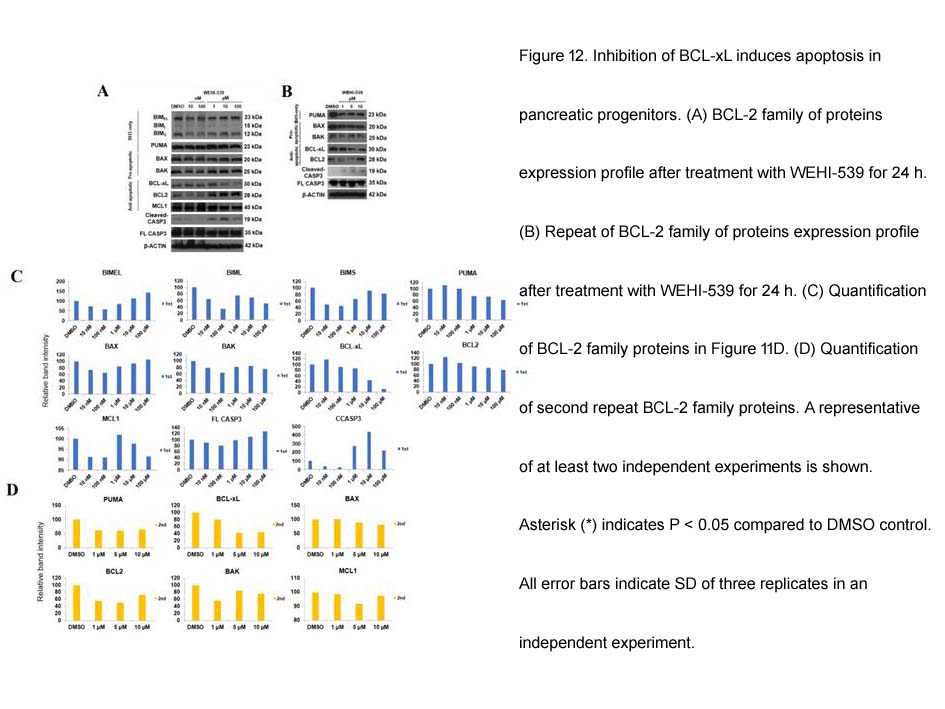
- 3. Loo, Larry Sai Weng. "Dynamics of BCL-2 family of proteins in early pancreatic progenitors and β-cells." Nanyang Technological University.
- 4. Boccellato, Chiara, et al. "Overcoming glioblastoma intractability: pre-clinical characterisation of TRAIL sensitisation by marizomib and novel treatment perspectives." Universität Stuttgart. 2022.
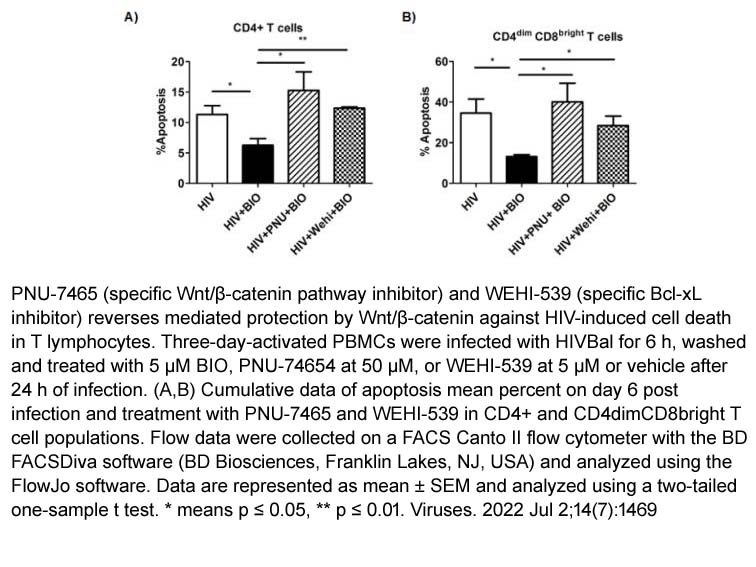
- 5. Yasmeen A. Albalawi, Srinivas D. Narasipura, et al. "Wnt/β-Catenin Protects Lymphocytes from HIV-Mediated Apoptosis via Induction of Bcl-xL." Viruses. 2022 Jul 2;14(7):1469. PMID: 35891449
- 6. Nadine Pollak, Aline Lindner, et al. "Cell cycle progression and transmitotic apoptosis resistance promote escape from extrinsic apoptosis." J Cell Sci. 2021 Dec 15;134(24):jcs258966. PMID: 34806752
- 7. Kirsteen J. Campbell, Susan M. Mason, et al. "Breast cancer dependence on MCL-1 is due to its canonical anti-apoptotic function." Cell Death Differ. 2021 Sep;28(9):2589-2600. PMID: 33785871
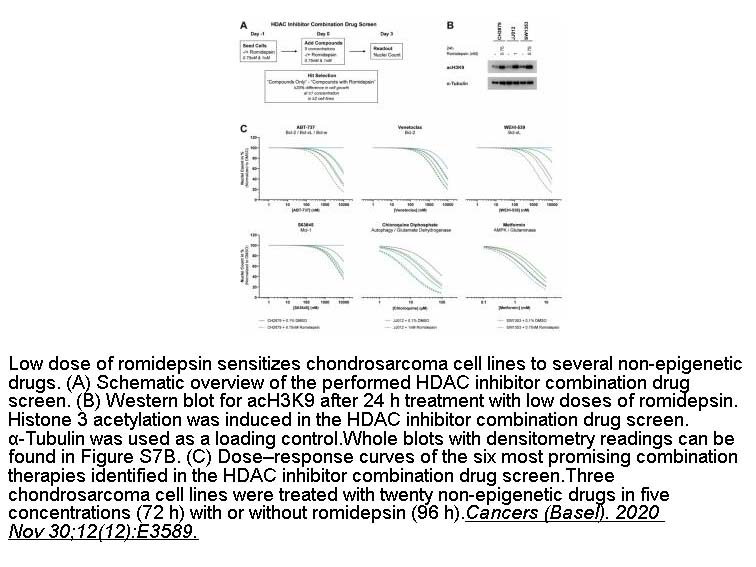
- 8. Sanne Venneker, Alwine B. Kruisselbrink, et al. "Beyond the influence of idh mutations: Exploring epigenetic vulnerabilities in chondrosarcoma." Cancers (Basel). 2020 Nov 30;12(12):E3589. PMID: 33266275
- 9. Enyuan Shang, Trang T. T. Nguyen, et al. "Epigenetic Targeting of Mcl-1 Is Synthetically Lethal with Bcl-xL/Bcl-2 Inhibition in Model Systems of Glioblastoma." Cancers 2020, 12(8), 2137;1 August 2020. PMID: 32752193
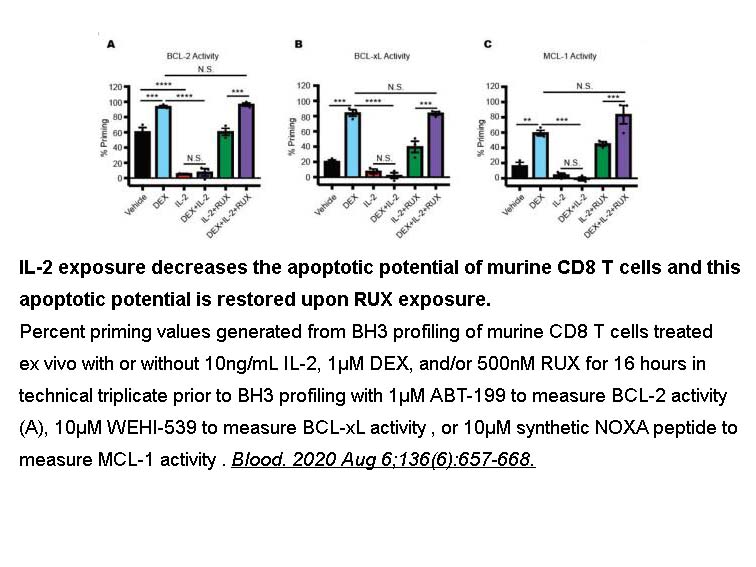
- 10. Meyer L, Verbist KC, et al. "JAK/STAT pathway inhibition sensitizes CD8 T cells to dexamethasone-induced apoptosis in hyperinflammation." Blood. 2020;blood.2020006075. PMID: 32530039
- 11. Loo LSW, Soetedjo AAP, et al. "BCL-xL/BCL2L1 is a critical anti-apoptotic protein that promotes the survival of differentiating pancreatic cells from human pluripotent stem cells." Cell Death Dis. 2020;11(5):378. PMID: 32424151
- 12. Villalobos-Ortiz M, Ryan J, et al. "BH3 profiling discriminates on-target small molecule BH3 mimetics from putative mimetics." Cell Death Differ. 2019 Jul 22. PMID: 31332296
- 13. de Jong Y, Monderer D, et al. "Bcl-xl as the most promising Bcl-2 family member in targeted treatment of chondrosarcoma." Oncogenesis. 2018 Sep 21;7(9):74. PMID: 30242253
- 14. Soderquist RS, Crawford L, et al. "Systematic mapping of BCL-2 gene dependencies in cancer reveals molecular determinants of BH3 mimetic sensitivity." Nat Commun. 2018 Aug 29;9(1):3513. PMID: 30158527
- 15. Wilson Xuan Mai."Comprehensive Characterization of the Apoptotic Machinery in Glioblastoma Identifies New Therapeutic Strategies." UNIVERSITY OF CALIFORNIA.2018-01-01.
- 16. Dai J, Luftig MA. "Intracellular BH3 Profiling Reveals Shifts in Antiapoptotic Dependency in Human B Cell Maturation and Mitogen-Stimulated Proliferation." J Immunol. 2018 Mar 1;200(5):1727-1736. PMID: 29358277
- 17. Vuillier C, Lohard S, et al. "E2F1 interacts with BCL-xL and regulates its subcellular localization dynamics to trigger cell death." EMBO Rep. 2017 Dec 12. PMID: 29233828
- 18. Mai WX, Gosa L, et al. "Cytoplasmic p53 couples oncogene-driven glucose metabolism to apoptosis and is a therapeutic target in glioblastoma." Nat Med. 2017 Nov;23(11):1342-1351. PMID: 29035366
- 19. Park HA, Licznerski P, et al. "Inhibition of Bcl-xL prevents pro-death actions of ΔN-Bcl-xL at the mitochondrial inner membrane during glutamate excitotoxicity."Cell Death Differ. 2017 Nov;24(11):1963-1974. PMID: 28777375
- 20. Tan Y, Lin Y, et al. "Selective Antagonism of Bcl-xL Potentiates M1 Oncolysis by Enhancing Mitochondrial Apoptosis." Hum Gene Ther. 2017 Jul 27. PMID: 28750564
- 21. Rose JC, Stephany JJ, et al. "Rapidly inducible Cas9 and DSB-ddPCR to probe editing kinetics." Nat Methods. 2017 Sep;14(9):891-896. PMID: 28737741
- 22. Russo M, Milito A, et al. "CK2 and PI3K are direct molecular targets of quercetin in chronic lymphocytic leukaemia." Oncotarget. 2017 Jun 27;8(26):42571-42587. PMID: 28489572
- 23. Bi C, Zhang X, et al."Inhibition of 4EBP phosphorylation mediates the cytotoxic effect of mechanistic target of rapamycin kinase inhibitors in aggressive B-cell lymphomas."Haematologica. 2017 Apr;102(4):755-764. PMID: 28104700
- 24. Pécot J, Maillet L, et al. "Tight Sequestration of BH3 Proteins by BCL-xL at Subcellular Membranes Contributes to Apoptotic Resistance." Cell Rep. 2016 Dec 20;17(12):3347-3358. PMID: 28009301
- 25. Bennett A, Sloss O, et al. "Inhibition of Bcl-xL sensitizes cells to mitotic blockers, but not mitotic drivers." Open Biol. 2016 Aug;6(8). PMID: 27512141
- 26. Kris Cameron Wood,Peter Saville Winter. "Compositions and Methods for Treating Cancer with JAK2 Activity." US Patent App. 15/027,216, 2016.
- 27. Winter PS, et al. "RAS signaling promotes resistance to JAK inhibitors by suppressing BAD-mediated apoptosis." Sci Signal. 2014 Dec 23. PMID: 25538080
| Physical Appearance | A solid |
| Storage | Store at -20°C |
| M.Wt | 583.72 |
| Cas No. | 1431866-33-9 |
| Formula | C31H29N5O3S2 |
| Synonyms | WEHI539,WEHI 539 |
| Solubility | insoluble in DMSO; insoluble in H2O; insoluble in EtOH |
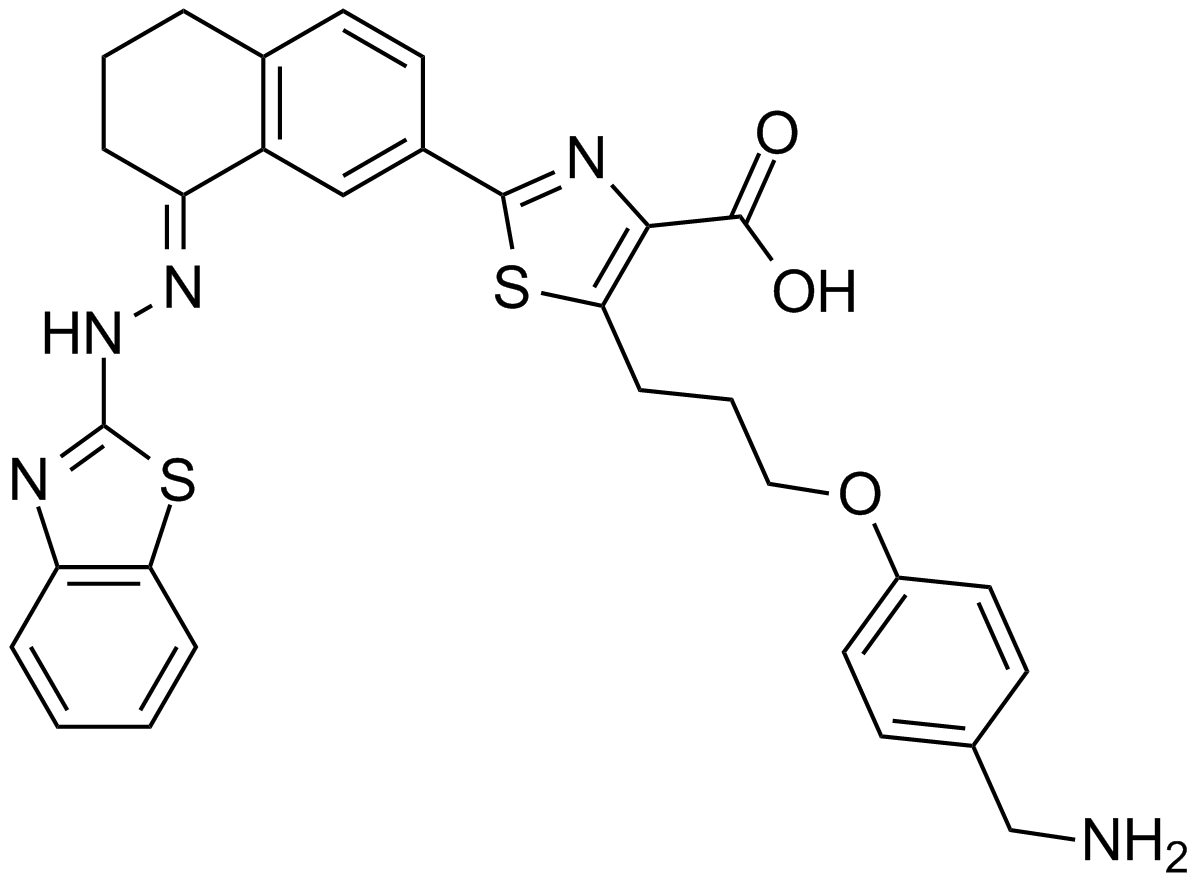
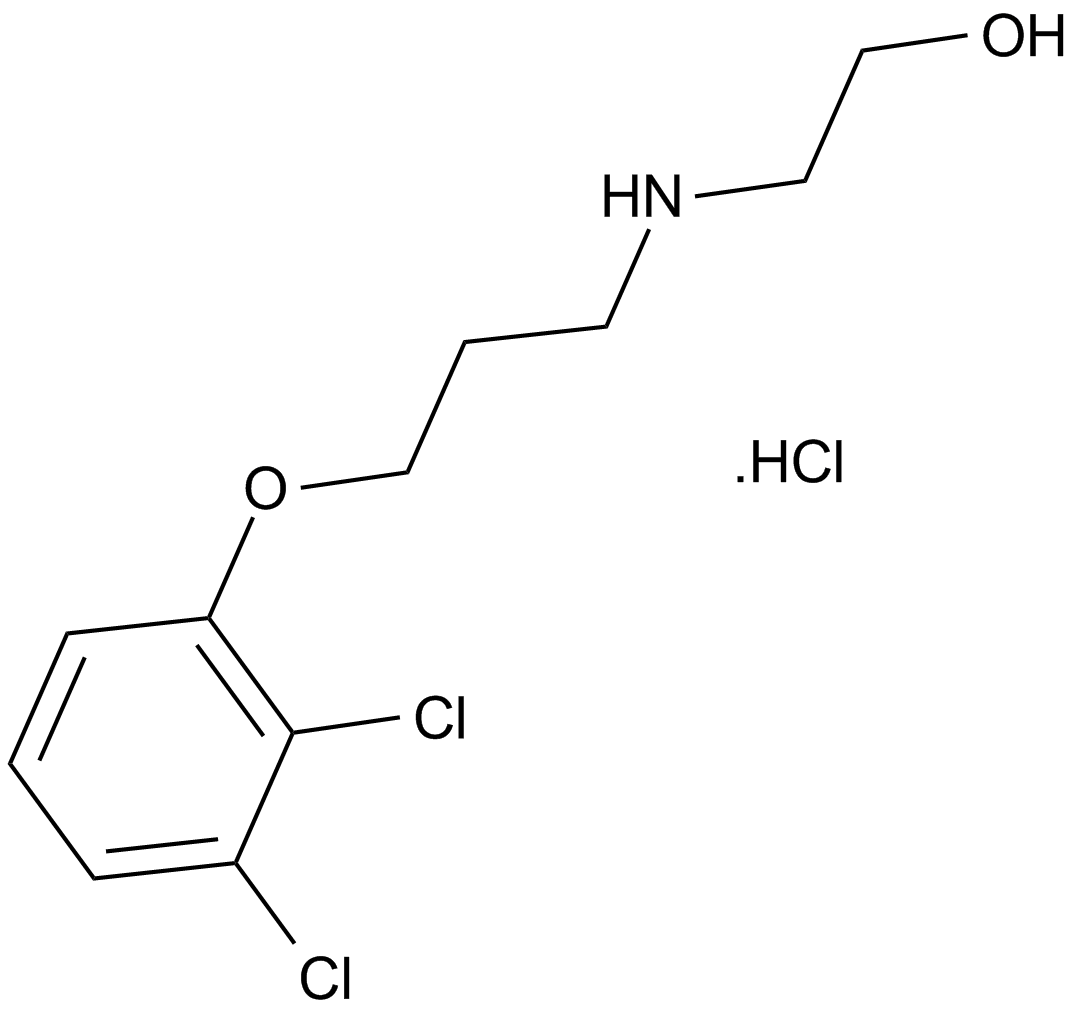
| Chemical Name | 5-[3-[4-(aminomethyl)phenoxy]propyl]-2-[(8E)-8-(1,3-benzothiazol-2-ylhydrazinylidene)-6,7-dihydro-5H-naphthalen-2-yl]-1,3-thiazole-4-carboxylic acid |
| SDF | Download SDF |
| Canonical SMILES | C1CC2=C(C=C(C=C2)C3=NC(=C(S3)CCCOC4=CC=C(C=C4)CN)C(=O)O)C(=NNC5=NC6=CC=CC=C6S5)C1 |
| Shipping Condition | Small Molecules with Blue Ice, Modified Nucleotides with Dry Ice. |
| General tips | We do not recommend long-term storage for the solution, please use it up soon. |
| Cell experiment[1]: | |
|
Cell lines |
Human colon cancer cell |
|
Reaction Conditions |
1 μM, 24h |
|
Applications |
Limiting dilution analysis with CSCs that were pre-treated with ABT-737, ABT-199 or WEHI-539 revealed that ABT-737 and WEHI-539 both were sufficient to decrease clonogenic capacity, whereas ABT-199 did not affect clonogenic growth. As WEHI-539 is selective for BCLXL, this points to a dependency of CSCs on BCLXL for survival. Importantly, ABT-737- or WEHI-539-induced loss of clonogenicity could be restored when BCLXL was ectopically overexpressed. When spheroid cultures were treated with ABT-737 or WEHI-539 compounds, CSCs were effectively sensitized toward oxaliplatin and other chemotherapeutic agents. |
|
References: 1. Colak S, Zimberlin CD, Fessler E et al. Decreased mitochondrial priming determines chemoresistance of colon cancer stem cells. Cell Death Differ. 2014 Jul;21(7):1170-7. |
|
| WEHI-539, has high affinity (subnanomolar) and selectivity for BCL-XL and potently kills cells by selectively antagonizing its prosurvival activity. WEHI-539 has a high affinity for BCL-XL (IC50 = 1.1 nM). | ||||||
| Targets | BCL-XL | |||||
| IC50 | 1.1 nM | |||||
Quality Control & MSDS
- View current batch:
Chemical structure
Related Biological Data
Related Biological Data
Related Biological Data
Related Biological Data
Related Biological Data
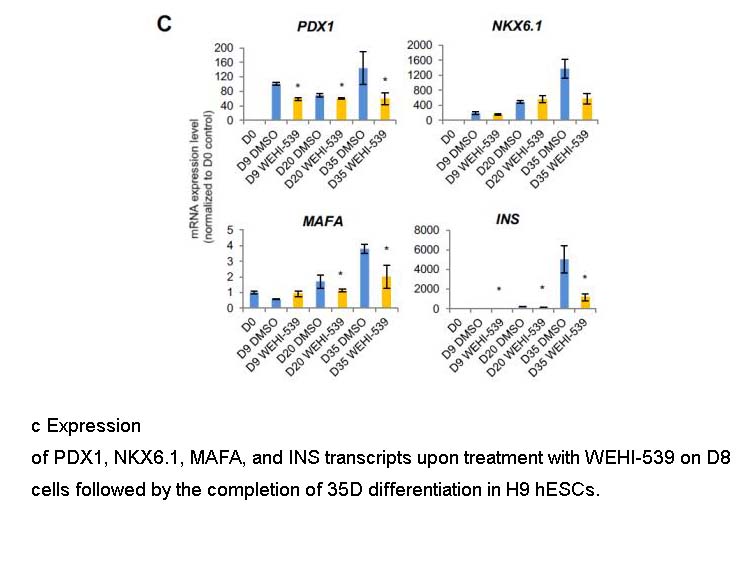
Related Biological Data
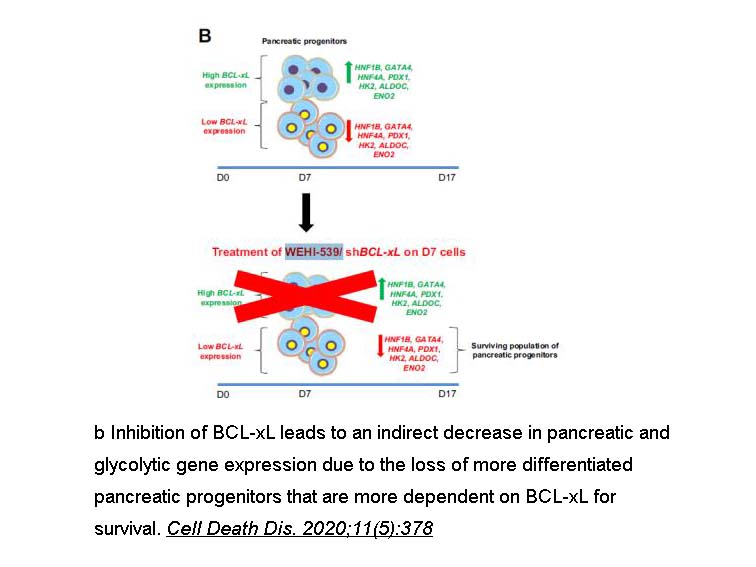
Related Biological Data