L189
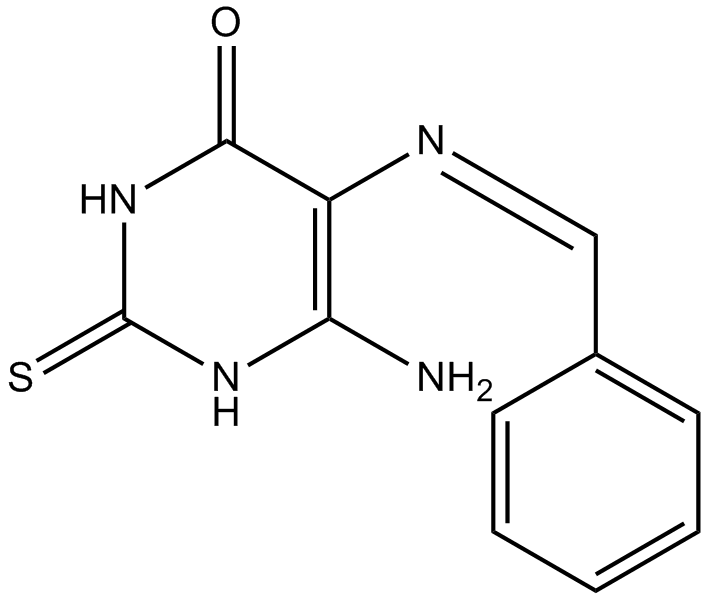
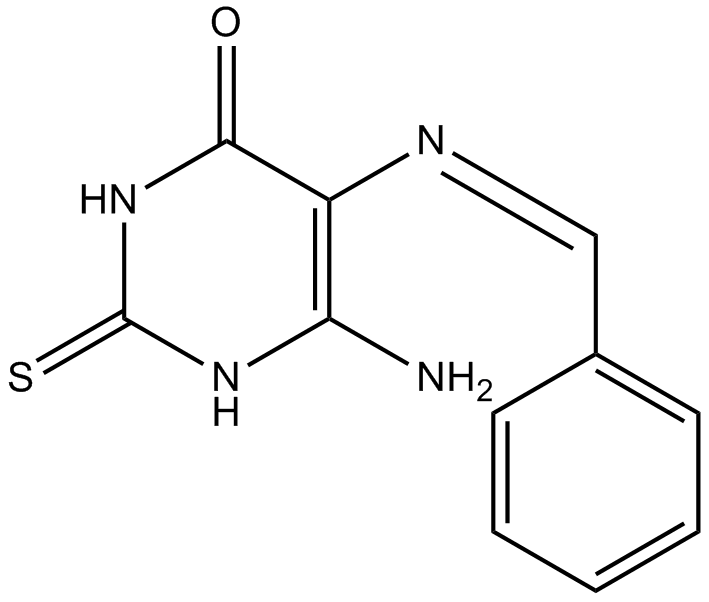
The chemical structure of L189 had been shown [1]. Three distinct chemical steps are involved in the formation of a new phosphodiester bond in DNA: (1) enzyme adenylylation, (2) adenylyl transfer to DNA, and (3) nick sealing [2]. L189 preferentially inhibits the second step [1]. L189 showed no inhibition even at a concentration of 100 μM to T4 DNA ligase. IC50 values for L189 to inhibit human DNA ligases I, III and IV are 5 ± 2 µM, 9 ± 2 µM and 5 ± 2 µM, respectively [1].
DNA ligase I catalyzes the ligation of single-strand breaks to complete DNA replication and repair. DNA ligase III is required for mitochondrial DNA replication and repair. DNA ligase IV is specialized for repair of nuclear double-strand breaks and is required for nonhomologous end joining and V (D) J recombination [2].
In a concentration-dependent manner, L189 reduced the viability and/or proliferation of a normal breast epithelial cell line MCF10A and the cancer cell lines HeLa, MCF7 and HCT116 established from cervical, breast and colon cancers, respectively. L189 also increase the rate of killed cells in cancer cell lines, especially HCT116 colon cancer cell lines and other cancer cell lines by ionizing radiation. But L189 did not increase the rate of killed cells in the normal cell line [1].
In vivo data and clinical trials are not yet available [3].
References:
[1]. Xi Chen, Shijun Zhong, Xiao Zhu, et al. Rational Design of Human DNA Ligase Inhibitors that Target Cellular DNA Replication and Repair. Cancer Res., 2008, 68(9): 3169-3177.
[2]. Mark R. Taylor, John A. Conrad, Daniel Wahl, et al. Kinetic Mechanism of Human DNA Ligase I Reveals Magnesium-dependent Changes in the Rate-limiting Step That Compromise Ligation Efficiency. Journal of Biological Chemistry, 2011, 286(26): 23054-23062.
[3]. Christian Jekimovs, Emma Bolderson, Amila Suraweera, et al. Chemotherapeutic compounds targeting the DNA double-strand break repair pathways: the good, the bad, and the promising. Frontiers in Oncology, 2014, 4: Article 86.
| Physical Appearance | A solid |
| Storage | Store at -20°C |
| M.Wt | 246.29 |
| Cas No. | 64232-83-3 |
| Formula | C11H10N4OS |
| Solubility | ≥62.5 mg/mL in DMSO; insoluble in EtOH; insoluble in H2O |
| Chemical Name | (Z)-6-amino-5-(benzylideneamino)-2-thioxo-2,3-dihydropyrimidin-4(1H)-one |
| SDF | Download SDF |
| Canonical SMILES | NC(NC(NC1=O)=S)=C1/N=C\c1ccccc1 |
| Shipping Condition | Small Molecules with Blue Ice, Modified Nucleotides with Dry Ice. |
| General tips | We do not recommend long-term storage for the solution, please use it up soon. |
| Cell experiment [1]: | |
|
Cell lines |
Human breast epithelial MCF10A cells, human colon cancer HCT116 cells, human cervical cancer HeLa cells, and human breast cancer MCF7 cells. |
|
Preparation method |
The solubility of this compound in DMSO is >10 mM. General tips for obtaining a higher concentration: Please warm the tube at 37℃ for 10 minutes and/or shake it in the ultrasonic bath for a while. Stock solution can be stored below -20℃ for several months. |
|
Reacting condition |
0~50 µmol/L for 6 days |
|
Applications |
In a concentration-dependent manner, L189 (0~50 µmol/L) reduced the proliferation and/or viability of a normal breast epithelial cell line MCF10A and the cancer cell lines MCF7, HeLa, and HCT116 established from breast, cervical, and colon cancers, respectively. In colony-forming assays, L189 was cytotoxic. L189 also increase the rate of killed cells in cancer cell lines, especially HCT116 colon cancer cell line and other cancer cell lines by ionizing radiation. But L189 did not increase the rate of killed cells in the normal cell line. |
|
References: [1] Christian Jekimovs, Emma Bolderson, Amila Suraweera, et al. Chemotherapeutic compounds targeting the DNA double-strand break repair pathways: the good, the bad, and the promising. Frontiers in Oncology, 2014, 4: Article 86. | |
Quality Control & MSDS
- View current batch:
Chemical structure