EdU Flow Cytometry Assay Kits (Cy5)
Measuring cell proliferation and cell cycle are a fundamental method to assess cell health, determine genotoxicity, and evaluate drug’s pharmacodynamic effect. The common method is measuring DNA synthesis directly. In previous experiments, there are several approaches such as the incorporation of radioactive nucleosides (3H-thymidine) or BrdU. Here, we introduce one new method, click chemistry-CuAAC (Copper-Catalyzed Azide-Alkyne Cycloaddition), and the use of this reaction in direct measurement of S-phase DNA synthesis in cell cycle.
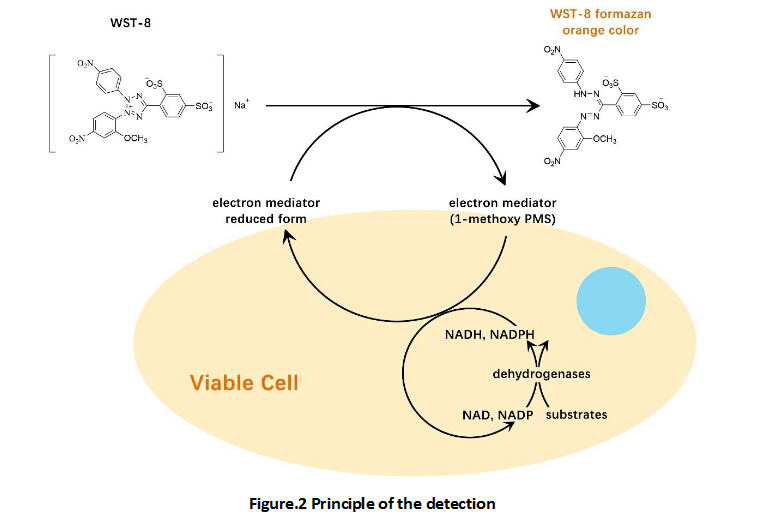
A nucleoside analog of thymidine, EdU (5-ethynyl-2’-deoxyuridine), can be incorporated into DNA strand during DNA synthesis. The alkynyl group of EdU is a biologically inert group that will undergo an extremely selective reaction with dye’s azido via a CuAAC reaction to afford an 1,2,3-triazole product. EdU and Cy5 azide possess biologically unique moieties to label DNA of proliferating cells, producing low backgrounds and high detection sensitivities. This CuAAC reaction affords superior regioselectivity and quantitative transformation under extremely mild conditions.
EdU Flow Cytometry Assay Kits (Cy5) is a sensitive and reliable way to detect and quantify cell proliferation in cells. It is optimized for flow cytometry.

- 1. Kentaro Takahashi, Julia Drolet, et al. "Diet-induced obesity induces oxidative stress and enhances H3K4me3 levels, driving nonresolving inflammation and myelopoiesis in hematopoietic stem and progenitor cells." J Immunol. 2025 Aug 7:vkaf156 PMID: 40795290
- 2. Fu-Gang Xiao, Zhou Yang, Shi-Yan Yu. "N7-methylguanosine-related gene decapping scavenger enzymes as a novel biomarker regulating epithelial cell function in diabetic foot ulcers." World J Diabetes. Nov 15, 2025; 16(11): 109455
- 3. Lan-Yue Ma, Zhao-Hua Deng, Ke Bai. "A single-cell hematopoietic microenvironmental atlas reveals progressive maturation of bone marrow vascular niche." Cell Regen. 2025 Dec 4;14(1):50 PMID: 41339619
- 4. Jason S. Wasserman, Bulat Faezov, et al. "FAM122A ensures cell cycle interphase progression and checkpoint control by inhibiting B55α/PP2A through helical motifs." Nat Commun. 2024 Jul 10;15(1):5776 PMID: 38982062
| Components | K1078-100 Test |
| EdU (Component A) | 20 mg |
| Cy5 Azide (Component B) | 1 vial |
| DMSO (Component C) | 8.5 mL |
| CuSO4 (100 mM Aqueous Solution) (Component D) | 1 mL |
| EdU Buffer Additive (Component E) | 400 mg |
Store the kit at -20ºC away from light and moisture, stable for 1 year. | |
Simple works in less time with no denaturation steps or harsh treatment
High efficiency with robust brightness
Compatible with some antibodies and cell cycle dyes
High consistency because of independence of variable anti-BrdU antibody